Our Research Teams and Our Director
Our Mission
The Center will be an innovative and sustainable scientific force in the creation of chemical probes, drug candidates, assay technologies, and computational approaches to advance understanding of biological mechanisms and place UNC at the forefront of translational medicine.

Kenneth H. Pearce, Ph.D.
Professor
Director, Center for Integrated Chemical Biology and Drug Discovery
Ken Pearce is a Professor and Director of the Center of Chemical Biology and Drug Discovery at the University of North Carolina at Chapel Hill. Pearce is also the Director of the Lead Discovery and Characterization group. His primary interest and expertise is the application of various methods for conducting early drug and chemical probe discovery research. These techniques include the development of biochemical and cell assays, high-throughput screening, DNA-encoded library technology, peptide phage display, and mechanistic/biophysical studies. Projects are typically collaborative efforts with labs across the UNC campus and span numerous therapeutic areas, with a particular focus on oncology. He is motivated by collaborative team-based and therapeutically-aligned science, training the next generation of discovery scientists, and contributing to service opportunities at UNC.

Samantha Pattenden, Ph.D.
Director, Applied Epigenetic Screening Technologies
ACCEPTING DOCTORAL STUDENTS
Lab Members
The Pattenden Lab develops innovative techniques in chromatin-based therapeutic target discovery and cancer diagnostics. Our research program enables discovery of novel molecular targets, pathways and mechanisms. Some of our current projects include:
Development of a cavitation enhancement technology to access archived tissues for epigenetics-based biomedical research
Collaborators: Dr. Ian Davis (Genetics, Pediatric Hematology/Oncology) and Dr. Paul Dayton (Biomedical Engineering)
Innovative new technologies that enable experimentally robust interrogation of epigenetic mechanisms are needed to broaden our understanding of epigenetic regulatory pathways in human development, disease, and therapeutic resistance. Formalin fixed, paraffin embedded (FFPE) tissues contain a wealth of information on human disease, however, extraction of high-quality chromatin (DNA together with associated nuclear proteins) from these samples for use in epigenetic assays has proven virtually impossible. We are exploring the use of a unique cavitation enhancement reagent in simplifying and standardizing chromatin extraction from FFPE tissues, with the goal of making archived biospecimens available for a broad range of epigenetic-based biomedical research.
Development of a First-in-Class High Throughput Assay Based on Chromatin Accessibility
Collaborators: Dr. Ian Davis (Genetics, Pediatric Hematology/Oncology
As appreciation of the importance of chromatin dysregulation in cancers has grown, chromatin regulatory proteins have emerged as promising targets for therapeutic discovery. Unlike alterations to the underlying DNA sequence, changes in chromatin states are both dynamic and reversible. Chromatin-related proteins are challenging targets for small molecule discovery since they frequently associate with multiple complexes with divergent cellular functions. For this reason, hit compounds derived from in vitro screening approaches based on a single chromatin-associated protein domain will likely suffer from significant off-target effects when tested in cells or in vivo.
To address the immediate need for cell-based chromatin screens, we have developed a target agnostic small molecule screening technology that exploits tumor-specific chromatin accessibility states as a relevant and direct functional readout. By targeting chromatin accessibility instead of a single transcription factor or chromatin regulatory protein, the limitations associated with in vitro screening and single protein target identification can be overcome. This screening technique represents a promising new paradigm in oncology drug discovery that will increase our mechanistic knowledge of tumor-specific changes in the epigenome and expand the repertoire of druggable targets.
Targeting ALT-cancer
Collaborators: Dr. Michael Jarster (Chemical Biology and Medicinal Chemistry
C ancerous cells develop a number of mechanisms to evade the checks and balances present in normal cells. One of the mechanisms by which tumor cells become immortal is by maintaining the length of the caps at the ends of chromosomes called telomeres. Telomere length is maintained by telomerase in ~85% of cancers and by the alternative lengthening of telomeres (ALT) pathway in the remaining ~15% of cancers. The ALT pathway is associated with poor prognosis, but the mechanisms by which ALT is induced are not well understood. ALT cells are frequently associated with the formation of extrachromosomal C-circles, which can be used as a readout for ALT activity. We are developing a high throughput assay using targeted small molecule chemical libraries to discover inhibitors of the ALT C-circle phenotype. Hit compounds will inform the key protein players in ALT maintenance and possibly reveal new therapeutic targets for ALT positive cancers.
ancerous cells develop a number of mechanisms to evade the checks and balances present in normal cells. One of the mechanisms by which tumor cells become immortal is by maintaining the length of the caps at the ends of chromosomes called telomeres. Telomere length is maintained by telomerase in ~85% of cancers and by the alternative lengthening of telomeres (ALT) pathway in the remaining ~15% of cancers. The ALT pathway is associated with poor prognosis, but the mechanisms by which ALT is induced are not well understood. ALT cells are frequently associated with the formation of extrachromosomal C-circles, which can be used as a readout for ALT activity. We are developing a high throughput assay using targeted small molecule chemical libraries to discover inhibitors of the ALT C-circle phenotype. Hit compounds will inform the key protein players in ALT maintenance and possibly reveal new therapeutic targets for ALT positive cancers.

Lindsey James, Ph.D.
Director, Chemical Biology
ACCEPTING DOCTORAL STUDENTS
Lab Members
 The goal of the James lab is to undertake and lead innovative and novel projects focused on the chemical biology of chromatin regulation, with an emphasis on the development of small molecule chemical probes. Providing such tool compounds to the scientific community has the potential to open new avenues of research in various disease relevant fields and translate to compounds of therapeutic value. Specifically, we are focused on developing compounds to study the domains that recognize the post-translational modification, methylated lysine. Aberrant methylation levels and ensuing changes in gene expression patterns resulting from the altered expression of methyl-lysine (Kme) regulatory proteins is one mechanism by which such epigenetic factors contribute to disease. Kme reader domains have emerged as less precedented epigenetic targets, yet considering the abundant links to cancer genetics, they are well suited to become the next impactful target class of chromatin regulators for intervention.
The goal of the James lab is to undertake and lead innovative and novel projects focused on the chemical biology of chromatin regulation, with an emphasis on the development of small molecule chemical probes. Providing such tool compounds to the scientific community has the potential to open new avenues of research in various disease relevant fields and translate to compounds of therapeutic value. Specifically, we are focused on developing compounds to study the domains that recognize the post-translational modification, methylated lysine. Aberrant methylation levels and ensuing changes in gene expression patterns resulting from the altered expression of methyl-lysine (Kme) regulatory proteins is one mechanism by which such epigenetic factors contribute to disease. Kme reader domains have emerged as less precedented epigenetic targets, yet considering the abundant links to cancer genetics, they are well suited to become the next impactful target class of chromatin regulators for intervention.
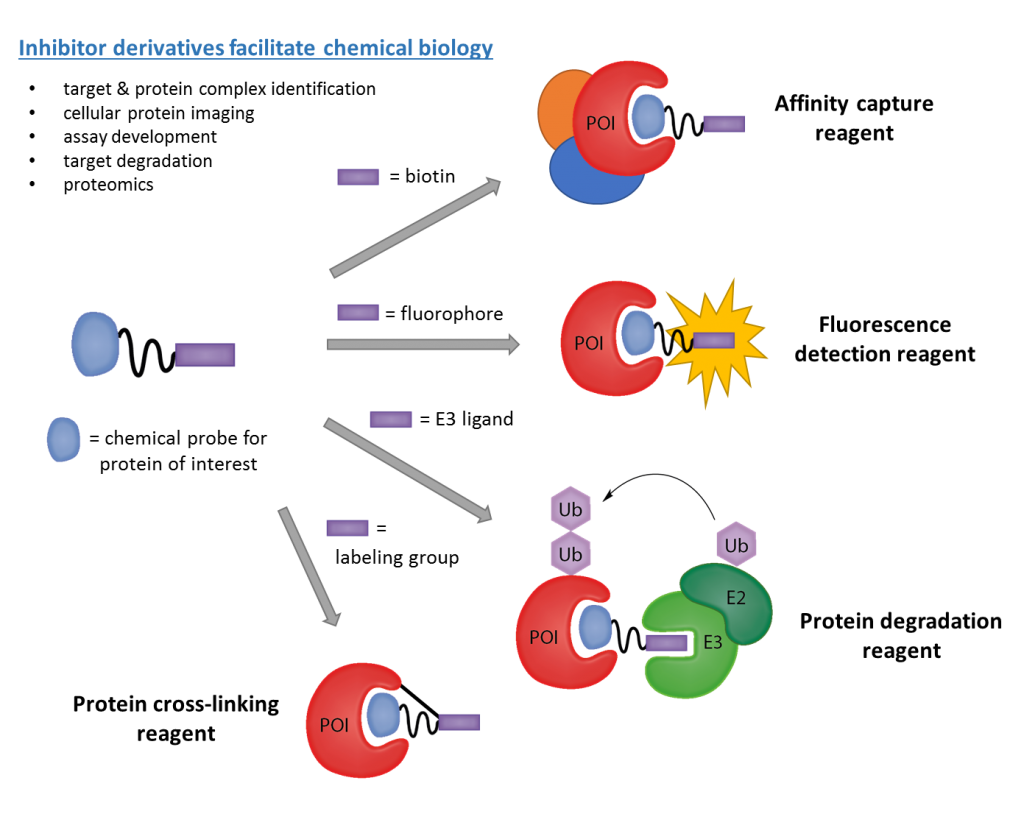
Our work in this area has pioneered the biochemical assays and medicinal chemistry strategies for high-quality probe development for the methyl-lysine reader target class, as well as the means by which to evaluate probe selectivity, mechanism of action, and cellular activity. Using a variety of approaches, we utilize such chemical tools to improve our understanding of their molecular targets and the broader biological consequences of modulating these targets in disease, particularly cancer. We also think about developing novel methods and screening platforms to discover hit compounds t o accelerate Kme reader probe discovery such as affinity-based combinatorial strategies, as well as novel ligand-based tools (see below) such as protein degradation reagents, or PROTACs, as potential therapeutic modalities. We have successfully developed numerous first-in-class chemical probes and currently have compounds in preclinical development (for example, see https://pharmacy.unc.edu/2019/12/pinnacle-hill-announces-first-project-agreement/).
o accelerate Kme reader probe discovery such as affinity-based combinatorial strategies, as well as novel ligand-based tools (see below) such as protein degradation reagents, or PROTACs, as potential therapeutic modalities. We have successfully developed numerous first-in-class chemical probes and currently have compounds in preclinical development (for example, see https://pharmacy.unc.edu/2019/12/pinnacle-hill-announces-first-project-agreement/).
Additionally, we collaborate closely with UNC faculty from various disciplines to provide medicinal chemistry and chemical biology expertise to bear on biological targets of therapeutic relevance.
Dr, James is committed to providing an inclusive, safe, and supportive research environment for trainees, teaching trainees to conduct rigorous, ethically sound, and responsible scientific research, and fulfilling the needs of trainees so that they can transition to the next phase of their careers.

Konstantin Popov, PhD
Director, Computational Biophysics
ACCEPTING DOCTORAL STUDENTS
Computational Biophysics (develops and applies computational tools to advance our understanding of complex biological systems and discover therapies for unmet medical needs.)
Dr. Konstantin Popov, Associate Professor in the Division of Chemical Biology and Medicinal Chemistry of the Eshelman School of Pharmacy directs our Computational Biophysics group here in the Center.
Konstantin comes to us from a Research Assistant Professor position in the Department of Biochemistry and Biophysics at UNC. His research expertise is at the interface of computational and structure-guided chemical probe and drug discovery, which is a vital part of the mission of the CICBDD.
Lab MembersResearch
The Popov Lab develops inventive, cutting-edge approaches to solve problems in modern computational structural biology and drug discovery. Our computational research, in collaboration with experimental screening and medicinal chemistry efforts in the Center, enables the identification of novel chemical probes and drug candidates to advance understanding of biological processes. Some of our recent projects include:
Identification and characterization of allosteric and cryptic binding sites
Allosteric protein regulation, achieved through chemical probes/modulators designed to bind to allosteric sites (usually distant from traditional orthosteric sites), enables effective control over protein functional activity. The potential structural diversity of allosteric sites allows for the design of chemical probes with high specificity and selectivity, minimizing potential future toxicity. Therefore, the identification of potentially allosteric binding sites and accessing their druggability is a promising tactic in modern drug discovery. We are developing and applying molecular dynamics (MD)-based, graph-theoretical, and AI-based approaches to study protein allostery in the context of drug discovery.
Development of AI-driven methods for accelerated virtual screening (VS)
The rapid expansion of purchasable, on-demand chemical libraries containing billions or even trillions of molecules has greatly advanced modern drug discovery. However, it has also posed significant challenges to the efficient application of traditional structure-based virtual screening methods. To address this challenge, we have been developing a novel computational methodology that greatly accelerates virtual screening. Our workflow uniquely integrates machine learning, generative chemistry, massive chemical similarity searching methods, and minimal molecular docking to nominate a small number of top-scoring virtual hits selected from ultra-large chemical libraries (e.g., 36B Enamine REAL Space). We have observed that our methodology yields the highest enrichment factors over random compound selection compared to state-of-the-art accelerated virtual screening methods, while requiring the least computational resources.
AI approaches for DNA-encoded library (DEL)-guided virtual screening (VS) and new-generation DEL library designs
DEL technologies have become increasingly popular as an effective hit and lead generation strategy, substantially reducing the cost per compound tested compared to traditional high-throughput screening. However, the method has several major limitations, including reliance on basic computational and visualization tools for manual data processing and analysis, off-DNA hit resynthesis for further confirmatory screenings and hit prioritization, and efficient combinatorial library design. These constraints can significantly delay the discovery of viable hits and leads. Thus, we are developing and implementing a comprehensive computational approach, in collaboration with Dr. Pearce’s lab, that utilizes DEL screening data and state-of-the-art AI modeling. This approach aims to identify and prioritize diverse, high-quality hits from commercially available compound libraries, as well as de novo generated chemical matter. Additionally, this methodology will help lay the foundation for new strategies to efficiently design target-specific DEL libraries, where AI models can suggest DEL fragments, purchasable or de novo, amenable to certain chemical reactions and more likely to have affinity toward specific families/classes of proteins.
Collaborative hit discovery and optimization projects
By utilizing both our in-house developed computational tools and publicly available resources, we make significant contributions to collaborative translational projects within the CICBDD. As the center we are dedicated to expanding our knowledge of biological targets and exploring the broader biological consequences that arise from modulating these targets.

Ken Pearce, Ph.D., Director
Director, Lead Discovery and Characterization
Lab Members
Currently, within the CICBDD Lead Discovery and Characterization Team, we develop comprehensive early drug discovery programs using a variety of methods including protein expression and purification, biochemical and cell assays, and screening in partnership with numerous collaborators at UNC. We have the ability to conduct compound screening using a library of over 200,000 drug-like small molecules assembled within the center. Our expertise is also in hit validation and triage methods, including the use of biophysical methods. Following successful hit discovery and validation, we work with collaborators and chemists within the center to produce structure-activity data and additional mode-of-action studies. Our ultimate goals are to: 1) produce high-quality chemical probes for exploring biology or drug targets, and 2) discover drug candidate molecules for the treatment of unmet medical needs.
The majority of our activities and interests are exploring targets within the epigenetic and  chromatin modification network and prosecuting novel targets, including signal transduction proteins, enzymes, and protein-protein interactions, for the discovery of new potential cancer treatments. These techniques include the development of biochemical and cell assays, high-throughput screening, DNA-encoded library technology, peptide phage display, and mechanistic/biophysical studies. Targets for these discovery campaigns include proteins involved in epigenetic regulation, signal transduction, and enzymatic modifications.
chromatin modification network and prosecuting novel targets, including signal transduction proteins, enzymes, and protein-protein interactions, for the discovery of new potential cancer treatments. These techniques include the development of biochemical and cell assays, high-throughput screening, DNA-encoded library technology, peptide phage display, and mechanistic/biophysical studies. Targets for these discovery campaigns include proteins involved in epigenetic regulation, signal transduction, and enzymatic modifications.

Xiaodong Wang, Ph.D.
Director, Medicinal Chemistry
Lab Members
The Wang lab is interested in developing drug leads/candidates for kinase, phosphate kinase and protein targets identified by UNC faculty and external investigators. We have successfully used the structure- and/or ligand-based drug design approaches to deliver compounds to clinic (MerTK  inhibitors such as MRX-2843) or licensing (IDH1 inhibitor, co-developed with NCATs). We will continue to apply similar approaches for drug discovery towards new targets.
inhibitors such as MRX-2843) or licensing (IDH1 inhibitor, co-developed with NCATs). We will continue to apply similar approaches for drug discovery towards new targets.
Our medicinal chemistry efforts are project-based with a faculty generated lead compound. We can support 3-5 hit to probe/lead projects concurrently. The typical timeline from completion of a screen to completion of leads is 1-2 years. Projects are prioritized primarily based on the quality of hits and project funding. To date, our most successful project has been the development of inhibitors of Mer kinase for the treatment of leukemia and immunosuppressive tumors, which has delivered a candidate MRX-2843 in clinical trials. A significant portion of this work was funded through a research contract as a member of NCI’s Chemical Biology Consortium (CBC). In addition, we were funded through the CBC to conduct structure activity relationship (SAR) studies to support lead development of inhibitors of IDH1 to treat glioblastoma. We have contributed to numerous drug discovery efforts with UNC faculty and external investigators. Most recently, we have been funded by NIH to develop probes for new targets to treat Alzheimer’s disease and MerTK/Axl dual and Tyro3 selective inhibitors for the treatment of cancer by targeting tumor cells and activating anti-tumor immunity.

