Craig Lee Lab
Craig Lee Lab
Craig Lee, Pharm.D, Ph.D. is a professor in the UNC Eshelman School of Pharmacy’s Division of Pharmacotherapy and Experimental Therapeutics (DPET). A key aspect of our mission is to optimize drug therapy through translating experimental and clinical pharmacology discoveries into precision medicine and accelerating application of these discoveries to improve patient care.
Dr. Lee is trained as a clinical/translational pharmaceutical scientist with expertise in cytochrome P450 metabolism, cardiovascular experimental therapeutics, and precision medicine/pharmacogenomics. He is an active member of the UNC McAllister Heart Institute and the UNC Program for Precision Medicine in Healthcare, and has an adjunct faculty appointment in the UNC School of MEdicine’s Division of Cardiology.
The overall objective of Dr. Lee’s research program is to improve the understanding of the central mechanisms underlying inter-individual variability in drug response as a means to develop novel therapeutic strategies that will improve public health.
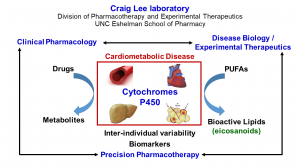
A major scientific focus of the Lee laboratory is the metabolism of drugs and eicosanoids by the cytochromes P450 enzyme system. The major therapeutic area of application of their research is cardiovascular and metabolic disease.
The Lee laboratory seeks to identify and elucidate the key factors that exacerbate inter-individual variability in the metabolism of and response to drugs currently on the market, and determine whether implementation of genomic and biomarker-guided drug selection and dosing strategies can reduce this variability in metabolism and response and improve patient outcomes.
The Lee laboratory also seeks to develop a thorough understanding of how cytochrome P450-derived eicosanoids (bioactive lipid mediators of arachidonic acid) regulate hepatic and extra-hepatic inflammatory responses, and determine whether modulation of this pathway will serve as an effective anti-inflammatory and end-organ protective therapeutic strategy for cardiovascular and metabolic disease.
Using genomics and biomarkers, the lab seeks to translate their preclinical discoveries into humans and determine which subsets of the population may be most likely to respond to the therapeutic strategies under evaluation in the laboratory.
The Lee laboratory is a highly collaborative and translational research program that integrates mechanistically-driven rodent and cell-based preclinical models with observational and interventional clinical studies. They have received funding from the National Institutes of Health and American Heart Association, authored over 100 manuscripts and over 100 abstracts in the areas of cytochromes P450, eicosanoid and drug metabolism, pharmacogenomics, and experimental therapeutics. Dr. Lee has served as the major research advisor for over 40 graduate students, post-doctoral fellows, and professional students.
Ongoing research projects in the Lee laboratory are highlighted below.
A major focus of Dr. Lee’s research program involves the implementation and evaluation of CYP2C19 genotype-guided selection of antiplatelet therapy in coronary artery disease patients. An algorithm that uses clinical factors and CYP2C19 genotype to improve precision of antiplatelet therapy selection in high-risk patients undergoing percutaneous coronary intervention was implemented at our institution in 2012. Dr. George A. Stouffer (UNC Chief of Cardiology; clinical implementation of the algorithm) and Dr. Lee (data collection and analysis) have collaborated to lead this ongoing effort. Data collection and analysis is focused on evaluating the impact of genotype and clinical factors on antiplatelet drug selection, and evaluating the impact of this strategy on clinical outcomes and cost. This ongoing project is funded by the NIH/NHLBI in collaboration with the University of Florida
Dr. Lee has been an affiliate member of the NIH Implementing GeNomics In PracTicE (IGNITE) Network since 2015, which seeks to enhance the use of genomic medicine by supporting the development of methods for incorporating genomic information into clinical care and exploration of the methods for effective implementation, diffusion and sustainability in diverse clinical settings. In collaboration with the IGNITE Pharmacogenomics Working Group, the UNC Program for Precision Medicine in Healthcare, and multiple DPET faculty, we are actively engaged in ongoing efforts to implement pharmacogenomics testing into clinical practice at our institution. Dr. Lee’s lab seeks to accomplish this in a manner that is evidence-based, feasible, sustainable, and amenable to the collection and analysis of data that enables continuous quality improvement, dissemination of best practices, and evaluation of impact on outcomes and cost.
Most drugs prescribed in pregnancy lack dosing information specific to this population. Dosing information is lacking, in part, because the key mechanistic factors that alter drug disposition (metabolism and transport) in pregnant people are poorly understood. This results in off-label prescribing and drug dosing, increased maternal and fetal toxicity, and reduced efficacy. Hepatic drug-metabolizing enzymes and drug transporters are integral to drug disposition. Consequently, key factors that influence hepatic drug metabolism and transport in pregnancy require systematic investigation.
Using a translational approach, ongoing projects in the Lee laboratory funded by the NIH/NICHD seek to elucidate how and to what extent pregnancy-related hormones alter hepatic metabolism and hepatobiliary disposition of clinically relevant drugs used in pregnancy that exhibit distinct clearance mechanisms in the liver. Their long-term goal is to predict pregnancy-evoked changes in hepatic drug disposition in vivo, define underlying mechanisms, and optimize drug selection and dosing in pregnant patients. A major area of the lab’s focus is on medications used to treat hypertensive disorders of pregnancy.
In parallel to the well-described cyclooxygenase and lipoxygenase pathways, cytochrome P450 (CYP) enzymes also synthesize biologically active eicosanoids in the cardiovascular system and constitute the “3rd pathway” of arachidonic acid metabolism. CYP-derived EETs and 20-HETE regulate numerous biological processes integral to the development and progression of cardiovascular and metabolic disease, such as blood pressure, inflammation, fibrosis, ischemia/reperfusion injury, angiogenesis, myocardial/vascular remodeling, and insulin resistance. Consequently, inhibitors of soluble epoxide hydrolase (sEH) and 20-HETE biosynthesis are in development.
Using a translational approach, the Lee lab seeks to obtain a deeper mechanistic understanding of these biological effects, in both preclinical models and in humans, in order to identify clinical applications for therapeutic strategies that modulate CYP-mediated eicosanoid metabolism and subsets of the population most likely to derive benefit from these experimental therapeutics.
Projects in the lab investigate the relationship between inter-individual variation in CYP-derived eicosanoids at the genetic and metabolite level and prognosis in human patients with atherosclerotic cardiovascular disease.
Craig Lee
Geri Messinger
Joyce Mosha
Daniel Sutton
Research Staff Alumni
- Alison Kannon, B.S. (2006-2011)
- Kimberly Vendrov, B.S. (2011-2015)
- Rebecca Rementer, M.S. (2016-2018)
- Natasha Kulick, B.A. (2018-2019, 2020-2021)
- Taryn Miner, M.S. (2020-2022)
- Grace Tillotson, B.S. (2021-2022)
- Joseph Grieco, B.S. (2022-2023)
Ashton Pearce
Danwei Shao
Madison Kalk
Research and Scholarship in Pharmacy pathway:
Current Students
- Layna Perini, Pharm.D. candidate (class of 2024)
RASP Alumni
- Melody Robbins, Pharm.D. (class of 2019)
- Jesse Martin, Pharm.D. (class of 2020)
- Rachel Black, Pharm.D. (class of 2020)
- Spenser Smith, Pharm.D. (class of 2020)
- Lindsay Ratner, Pharm.D. (class of 2022)
- Julian Garcia, Pharm.D. candidate (class of 2023)
- Valeria Laboy Collazo, Pharm.D. candidate (class of 2023)
- Marshall Winget, Pharm.D. candidate (class of 2023)
Honors Program Alumni
- Alexandra Cervantes, Pharm.D. (class of 2018)
- Nan Wang, Pharm.D. (class of 2017)
- Amy Li, Pharm.D. (class of 2017)
- Michael Wells, Pharm.D. (class of 2016)
Graduate Students
| Name | Year(s) | Degree(s) | Current Position |
| Katherine Theken | 2006-2011 | Pharm.D., Ph.D. | Assistant Professor, Oral Surgery/Pharmacology, School of Dental Medicine, University of Pennsylvania |
| Robert Schuck | 2008-2013 | Pharm.D., Ph.D. | Deputy Director, Division of Translational and Precision Medicine, Office of Clinical Pharmacology, U.S. Food and Drug Administration |
| Akin Oni-Orisan | 2010-2015 | Pharm.D., Ph.D. | Assistant Professor of Pharmacy, Department of Clinical Pharmacy, University of California-San Francisco |
Postdoctoral Research Associates
| Name | Year(s) | Degree(s) | Current Position |
| Yangmei Deng | 2007-2011 | M.D., Ph.D. | Research Specialist, UNC Marsico Lung Institute |
| Weibin Zha | 2012-2013 | B.S. Pharm.D., Ph.D. |
Director, DMPK Recursion |
| Raju Khatri | 2017-2019 | M.S., Ph.D., DABT | Manager, DMPK, SK Life Science |
| Tien Van Le | 2022-2023 | Pharm.D. | Graduate Student, Pharmaceutical Sciences, University of Pittsburgh |
Clinical Research / Drug Development / Medical Affairs Fellows
| Name | Year(s) | Degree(s) | Current Position |
| Almasa Bass | 2007-2008 | Pharm.D. | Clinical Program Director, UCB |
| Kyle Ellis | 2008-2009 | Pharm.D. | Senior Clinical Manager, Specialty Dispensing Operations, CVS Health |
| Bryant Tran | 2009-2010 | Pharm.D., M.S. | Scientific Director, Medical Affairs GlaxoSmithKline |
| Savanna Steele | 2010-2011 | Pharm.D. | Director, Feasibility Strategy, PPD |
| Brian P. Simmons | 2011-2012 | Pharm.D., MSCR | Associate Group Director, Clinical Science, Genentech |
| Robert Wittorf | 2012-2013 | Pharm.D. | Director, Quality Compliance, Merck |
| John Andrew Lee | 2013-2014 | Pharm.D. | Senior Medical Science Liaison, Immuno-Oncology, Bristol Meyers Squibb |
| Melissa Polasek | 2014-2015 | Pharm.D. | Senior Director, Clinical Development, Imugene Limited |
| Hitesh Patel | 2017-2018 | Pharm.D. | Director, Medical Affairs, GlaxoSmithKline |
| Alexis Williams | 2018-2019 | Pharm.D. | Senior Medical Reviewer, Indivior |
| Megan Gower | 2019-2020 | Pharm.D. | Clinical Scientist, Takeda |
| Ian Mulrenin | 2020-2021 | Pharm.D. | Associate Clinical Scientist, PPD |
| Anh Nguyen | 2021-2022 | Pharm.D. | Clinical Development Scientist, Oncology, Gilead Sciences |
| Kayla Tunehag | 2022-2023 | Pharm.D., MSPH | Clinical Research/Drug Development Fellow, PPD |









